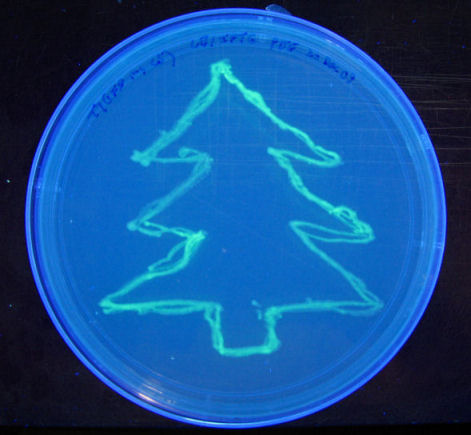
Product Review: BioTek Microflo liquid dispensing machine
A couple of pictures of the BioTek Microflo liquid dispenser.
Device with hatch open and spring tension unlocked:
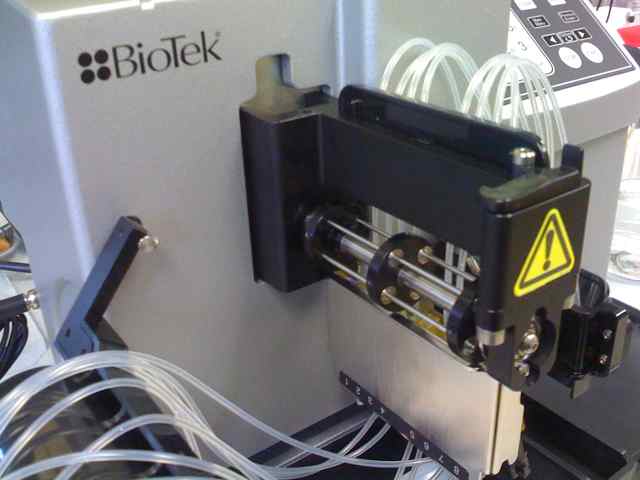

BioTek Microflow cartridge in place. Spring tensioner open. Eight silicone tubes run in parallel, across the pump axle.
Device with hatch open and spring tension locked:
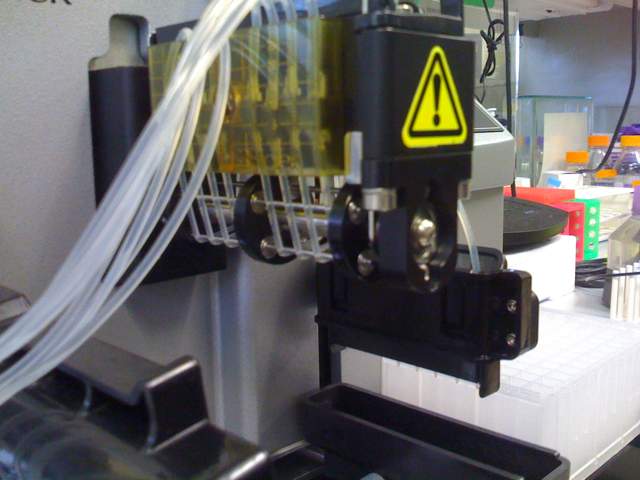

Biotek Microflo with tensioner locked. Tension is placed across the silicone tube liquid lines. Tension is adjustable with a set screw parallel to each line (screws not shown here, they are vertical and can be seen in a top-down view). The axle rotates clockwise or counterclockwise to move liquid forward or backward with peristaltic action. It is very fast.
This machine has both serial RS-232 and USB; however, the communication link is a proprietary protocol which is only compatible with BioTek’s Microsoft Windows software. The machine is not Unix compatible. So unfortunately, I won’t be using this device in my lab automation setup.
U.S. Office of Science and Technology Policy soliciting your feedback on “Improving Public Access to Results of Federally Funded Research” until Dec 20, 2009
The U.S. Office of Science and Technology Policy, under directives from the President Obama administration, is soliciting public feedback. Note the deadline! (Dec. 10th-20th)
Policy Forum on Public Access to Federally Funded Research: Implementation
Thursday, December 10th, 2009 at 7:25 pm by Public Interest Declassification Forum
By Diane DiEuliis and Robynn Sturm
Yesterday we announced the launch of the Public Access Forum, sponsored by the White House Office of Science and Technology Policy. Beginning with today’s post, we look forward to a productive online discussion.
One of our nation’s most important assets is the trove of data produced by federally funded scientists and published in scholarly journals. The question that this Forum will address is: To what extent and under what circumstances should such research articles—funded by taxpayers but with value added by scholarly publishers—be made freely available on the Internet?
The Forum is set to run through Jan. 7, 2010, during which time we will focus sequentially on three broad themes (you can access the full schedule here). In the first phase of this forum (Dec. 10th-20th) we want to focus on the topic of Implementation. Among the questions we’d like to have you, the public and various stakeholders, consider are:
- Who should enact public access policies? Many agencies fund research the results of which ultimately appear in scholarly journals. The National Institutes of Health requires that research funded by its grants be made available to the public online at no charge within 12 months after publication. Which other Federal agencies may be good candidates to adopt public access policies? Are there objective reasons why some should promulgate public access policies and others not? What criteria are appropriate to consider when an agency weighs the potential costs (including administrative and management burdens) and benefits of increased public access?
- How should a public access policy be designed?
- Timing. At what point in time should peer-reviewed papers be made public via a public access policy relative to the date a publisher releases the final version? Are there empirical data to support an optimal length of time? Different fields of science advance at different rates—a factor that can influence the short- and long-term value of new findings to scientists, publishers and others. Should the delay period be the same or vary across disciplines? If it should vary, what should be the minimum or maximum length of time between publication and public release for various disciplines? Should the delay period be the same or vary for levels of access (e.g. final peer reviewed manuscript or final published article, access under fair use versus alternative license)?
- Version. What version of the paper should be made public under a public access policy (e.g., the author’s peer-reviewed manuscript or the final published version)? What are the relative advantages and disadvantages of different versions of a scientific paper?
- Mandatory v. Voluntary. The NIH mandatory policy was enacted after a voluntary policy at the agency failed to generate high levels of participation. Are there other approaches to increasing participation that would have advantages over mandatory participation?
- Other. What other structural characteristics of a public access policy ought to be taken into account to best accommodate the needs and interests of authors, primary and secondary publishers, libraries, universities, the federal government, users of scientific literature and the public?
We invite your comments […]
Give government your feedback on how to release data and publications from publicly funded research.
- Register for an account on the U.S. Office of Science and Technology Policy site here: http://blog.ostp.gov/wp-login.php?action=register
- Leave your policy comments on their WordPress blog!
More information is in the U.S. Office of Science and Technology Policy video:
Microplate Standard Dimensions
The mission of the Microplate Standards Working Group (MSWG) is to recommend, develop, and maintain standards to facilitate automated processing of microplates on behalf of and for acceptance by the American National Standards Institute (ANSI). Once such standards are approved by the MSWG, they are presented to the governing council of the Society of Biomolecular Screening (SBS) for approval for submission to the ANSI. Although sponsored by SBS, membership in the MSWG is open to all interested parties directly and materially affected by the MSWG’s activities, including parties who are not members of SBS.
- ANSI/SBS 1-2004: Microplates – Footprint Dimensions
- ANSI/SBS 2-2004: Microplates – Height Dimensions
- ANSI/SBS 3-2004: Microplates – Bottom Outside Flange Dimensions
- ANSI/SBS 4-2004: Microplates – Well Positions
“Meat 2.0”
In synthetic biology conferences, engineering improvements of food is listed in the top three applications of the new technology. As an example, George Church’s lab developed a genetic engineering technology specifically aimed at evolving super-tomatoes containing high amounts of the anti-oxident lycopene, as proof-of-concept. Frequent brainstorming “what could syn bio do?” sessions include ideas of growing thick beef steaks without the cow: in essence, this is presumed to be an improvement on quality, cleanliness, nutrition, and animal rights, than today’s factory-farming method of bringing steak to the table.
What if there is already a better “steak”? Let’s call it Meat 2.0. How about modifying Rhizopus oligosporus, the fungus used in making tempeh, to create new tastes or additional vitamins? Note that the below article states, “cost of preparing 1.5 kg of tempeh was less than US$1.”
Nutritional and sensory evaluation of tempeh products made with soybean, ground-nut, and sunflower-seed combinations
M. P. Vaidehi, M. L. Annapurna, and N. R. Vishwanath
Department of Rural Home Science and Department of Agricultural Microbiology, University of Agricultural Sciences, Bangalore, IndiaINTRODUCTION
Tempeh products made from soybeans and from combinations of soybeans with ground-nuts and sunflower seed at ratios of 52:48 and 46:54 respectively were tested for their appearance, texture, aroma, flavour, and over-all acceptability. In addition, tempeh was prepared with and without the addition of bakla (Vicia faba) to soybeans in various ratios to obtain a tempeh of acceptable quality and nutritional value (1). Bakla tempeh at a 1:1 ratio was found to be crisper and more palatable than plain soybean tempeh, but at 3:1 the tempeh had a mushroom odour.
EXPERIMENTS
Materials
Tempeh culture (Rhizopus oligosporus) was obtained from the New Age Food Study Center, Lafayette, California, USA. It was grown on a rice medium and inoculated while different blended tempehs were prepared. A 2.5 9 packet of culture was used for 250 9 of substrate on a dry weight basis.
Soybeans (Hardee), ground-nuts (TMV-30), and sunflower seed (Mordon) were obtained from the University of Agricultural Sciences, Bangalore. Three varieties of tempeh -100 per cent soy, soy-ground-nut (52:48), and soy” sunflower seed (46:54)-were prepared under identical conditions.
Preparation of Tempeh and Products
“ELISA Redux” 96-Well Plate Cryptography Challenge
The publication GEN is running a contest, with $1,500 plus fancy biotechnology equipment as the prize, for the first one who can decode the cryptographic message hidden in this 96-well plate:
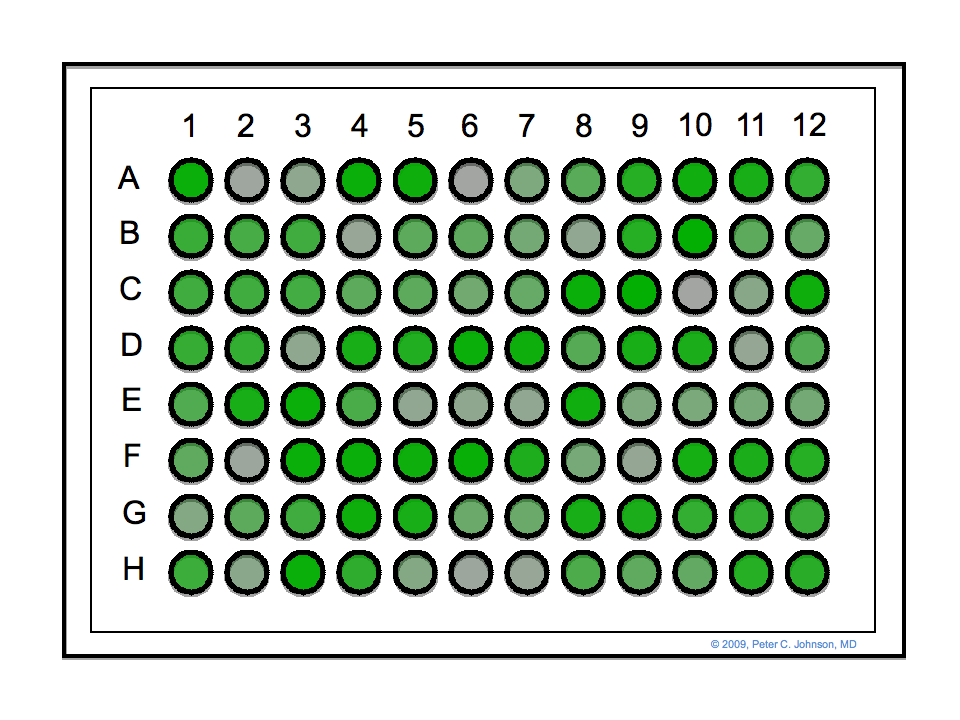
"A message has been encrypted into the ELISA plate image, called ELISA Redux, based on the color of each well."
Battery-powered, Pocket-sized PCR Thermocycler
A few years ago, some bright students at Texas A&M improved upon the most basic tool for manipulating microbiology: a thermocycler. Thermocyclers are typically large tabletop instruments which require a large sample, a lot of electrical power, and a lot of time to heat and cool. Alternatively, the process can be done by hand with a pot of boiling water, a bucket of cold water, a stopwatch, and a lot of free time and patience. The sample itself, for the purpose of “amplifying” the desired material in the sample, runs through many iterations of heat/cool cycling, such as the following:
- Denature: 95°C, 15 mins
Thermocycling
- No. of cycles: 39
- Denature: 94°C, 30 secs
- Anneal: 62°C, 30 secs
- Elongate: 68°C, 3.5 mins
Termination
- Elongate: 68°C, 20 mins
- Hold: 4°C, until removed from machine
The Texas team created a pocket-sized version, which could run on batteries as well, and most notably, was able to create a new patent. By creating a pocket-sized, battery powered device, the team accomplished several very important features:
- The device can be used easily at the point-of-care, as a field unit;
- The device is much lower engineering cost, and much lower patent royalty cost: from thousands of dollars down to hundreds of dollars;
- The device uses a much smaller sample, and has faster heat/cool times, thus reducing the experimental cost and experimental time.
This new thermocycler has created some excitement in the bio community for some time — however, it still took over a year to finish patent issues. The university owned the patent; they requested royalties, and up-front option fees, and meanwhile, the device itself remained in limbo. The great news is that the manufacturing prototype is announced (see below). The bad news is that such patent hassles are typical, and this was a simple case where a university owned the patent in whole rather than multiple holders owning the patent in part.
From the business angle, the thermocycler market is a billion-dollar market, since it is a fundamental tool for all microbiology or genetic engineering labs.
Some of original papers and articles for the “$5 thermocycler” are:
- Engaget: Mini DNA replicator costs $10 – May, 2007
- New Scientist: Mini DNA replicator could benefit world’s poor – May, 2007
- Angewandte Chemie: A Pocket-Sized Convective PCR Thermocycler – April, 2007
From Rob Carlson’s synthesis.cc blog:
This week Biodesic shipped an engineering prototype of the LavaAmp PCR thermocycler to Gahaga Biosciences. Joseph Jackson and Guido Nunez-Mujica will be showing it off on a road trip through California this week, starting this weekend at BilPil. The intended initial customers are hobbyists and schools. The price point for new LavaAmps should be well underneath the several thousand dollars charged for educational thermocyclers that use heater blocks powered by peltier chips.
The LavaAmp is based on the convective PCR thermocycler demonstrated by Agrawal et al, which has been licensed from Texas A&M University to Gahaga. Under contract from Gahaga, Biodesic reduced the material costs and power consumption of the device. We started by switching from the aluminum block heaters in the original device (expensive) to thin film heaters printed on plastic. A photo of the engineering prototype is below (inset shows a cell phone for scale). PCR reagents, as in the original demonstration, are contained in a PFTE loop slid over the heater core. Only one loop is shown for demonstration purposes, though clearly the capacity is much larger.
The existing prototype has three independently controllable heating zones that can reach 100C. The device can be powered either by a USB connection or an AC adapter (or batteries, if desired). The USB connection is primarily used for power, but is also used to program the temperature setpoints for each zone. The design is intended to accommodate additional measurement capability such as real-time fluorescence monitoring.
We searched hard for the right materials to form the heaters and thin film conductive inks are a definite win. They heat very quickly and have almost zero thermal mass. The prototype, for example, uses approximately 2W whereas the battery-operated device in the original publication used around 6W.
What we have produced is an engineering prototype to demonstrate materials and controls — the form factor will certainly be different in production. It may look something like a soda can, though I think we could probably fit the whole thing inside a 100ml centrifuge tube.
If I get my hands on one myself, I’ll post a review.
Add Streaming Video to any Bio-lab!
Combining an inexpensive (under $15) USB webcam with free VLC media player software, it is simple to add password-protected internet streaming video for remote users to any lab. VLC includes the ability to capture from a local webcam, transcode the video data, and stream the video over the web. It’s available for OS/X, Unix, Linux, and Microsoft systems.
Hint: Video formats are confusing. Even video professionals have a tricky time figuring out the standards and compatibility issues. Today’s web browsers also have limitations in what they can display (mime types and such) — which simply means both sides need to use VLC. Figuring all this out using the VLC documentation takes some work. Transcoding the video is required and a proper container must be used to encapsulate both video and audio. Once debugged, it’s good to go.
Here’s how it worked in the lab:
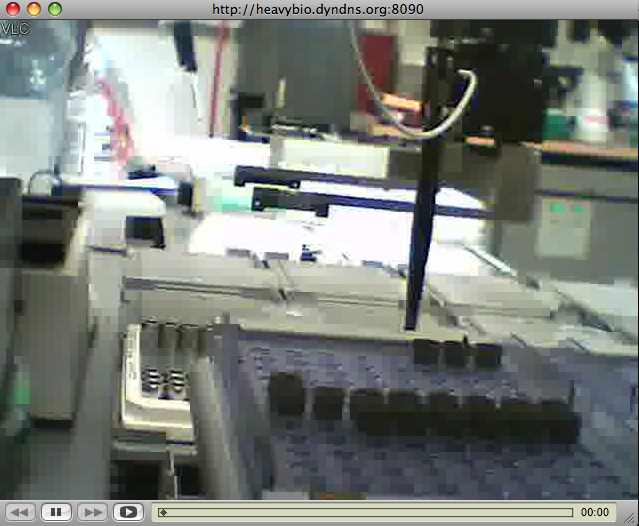
See the setup below to get it running.
More on Bio-lab Automation – Software for Controlling FIAlab Devices for Microfluidics
Perl software to control lab syringe pump and valve device, for biology automation, initial version finished today. Works great. Next, need to add the network code, it can be controlled remotely and in synchronization with other laboratory devices, including the bio-robot. This software will be used in the microfluidics project. The software is also part of the larger Perl Robotics project, and a new release will be posted to CPAN next week.

More details on the software follow:
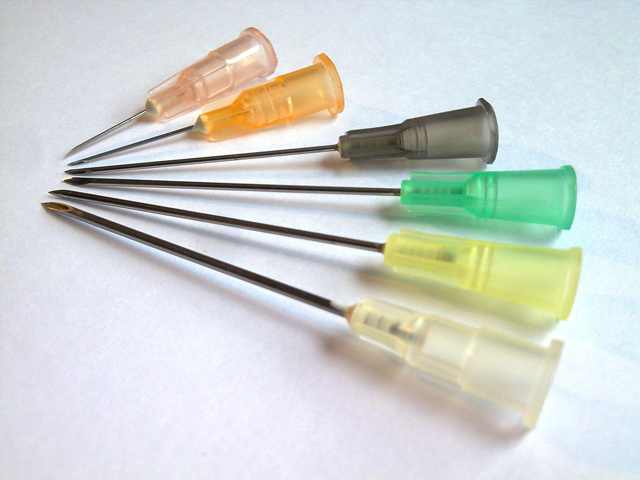
When a needle is not a needle: inside & outside needle diameter variations
In various discussions in biology circles there’s often the lament that “biology is hard” (which I agree) and from the biologists there are continued remarks that repeating a protocol in a slightly different way will have poor results. As an engineer I am fascinated by this because reproducibility is the key to making biology “easier to engineer”. Once a method is reproducible in different environments then it can be made into a black box for reuse without worrying about whether it will work under slightly varying conditions.
In one research paper, the chemical engineers dug into part of the reason why their experiment had differing results. They found that the size of needles varied considerably even within the same gauge and same vendor.

Don’t Always Trust Open Source Software. Why Trust Open Source Biology?
The software you are happily using may be.. unnecessarily brittle. Recently I’ve been developing a little bit of high-level software using open source libraries. Sometimes it amazes me that open source software works at all. Here’s an excerpt from the internals I found in the open source library when I looked at why it might not be working properly:
if (Pipe){
while(iFlag){
vpData = Pipe->Read(&dLen);
iFlag = 0;
// If we have more data to read then for God's sake, do it!
// I don't know if this will work ... it would return an
// array. This may not be good. Hmmmm.
if(!vpData && GetLastError() == ERROR_MORE_DATA){
iFlag = 1;
}
if(dLen){
XPUSHs(sv_2mortal(newSVpv((char *)vpData, dLen)));
}else{
sv_setsv(ST(0), (SV*) &PL_sv_undef);
}
}
The standard responses from the “open source rah-rah crowd” are something like the following:
- “Yeah, that’s a crazy comment, but at least you can see it! In proprietary software, there’s the same problems, it’s just hidden!”
- “At least you’re given the source code so you can fix it! In proprietary software, you’re never given access to the source code so you couldn’t fix it if you wanted to!”
These responses miss the big point that commercial software is often much more fully tested for it’s specific environment, and undergoes a much more rigorous design process. (Beyond the designed-for environment, things might break. However, the environment is usually described.)
Having something that works — even if it isn’t “great” software — is better than not having anything at all; so on the whole, we can’t complain too much. Open source is expected to evolve, over the long term (meaning, decades), into a better system: it’s assumed that eventually, most of the oddities will be ironed out. The Linux kernel itself contains similar comments (I’ve seen them in debugging the UDP/IP stack) which is astounding considering that non-professionals consider Linux to be “stable”. Kernel hackers know the truth — it “mostly” works (with “mostly” being better than “nothing”).. Next time someone offers you “free software” take a moment to think: how much do I have to trust that software to work in a situation which may be different than the author’s original working environment? How much of the code’s architecture might have comments such as “Hmm.. This isn’t supposed to work or might not work..”? How much is it going to cost ($$$) to find the oddities and dig into the internals to fix them?
The connection to Biology here is that these crazy design comments like “Hmm.. It really isn’t proper design to build it this way.. but it seems to be work” in synthetic life will be too small to ever read. (In the RNA or DNA.) At least with open source software, there’s a big anti-warrantee statement; don’t use the software if there is liability involved. As I posted last year, the “Open Biology License” hasn’t touched on liability issues at all — only patent issues. How much can Open Biology be trusted, how much might it cost ($$$) to dig in to find the strange biological behavior, and attempt to fix them? Debugging biology is much, much harder than debugging software.
Perl Bio-Robotics module, Robotics.pm and Robotics::Tecan
FYI for Bioperl developers:
I am developing a module for communication with biology robotics, as discussed recently on #bioperl, and I invite your comments. Currently this module talks to a Tecan genesis workstation robot. Other vendors are Beckman Biomek, Agilent, etc. No such modules exist anywhere on the ‘net with the exception of some visual basic and labview scripts which I have found. There are some computational biologists who program for robots via high level s/w, but these scripts are not distributed as OSS.
With Tecan, there is a datapipe interface for hardware communication, as an added $$ option from the vendor. I haven’t checked other vendors to see if they likewise have an open communication path for third party software. By allowing third-party communication, then naturally the next step is to create a socket client-server; especially as the robot vendor only support MS Win and using the local machine has typical Microsoft issues (like losing real time communication with the hardware due to GUI animation, bad operating system stability, no unix except cygwin, etc).
On Namespace:
I have chosen Robotics and Robotics::Tecan. (After discussion regarding the potential name of Bio::Robotics.) There are many s/w modules already called ‘robots’ (web spider robots, chat bots, www automate, etc) so I chose the longer name “robotics” to differentiate this module as manipulating real hardware. Robotics is the abstraction for generic robotics and Robotics::(vendor) is the manufacturer-specific implementation. Robot control is made more complex due to the very configurable nature of the work table (placement of equipment, type of equipment, type of attached arm, etc). The abstraction has to be careful not to generalize or assume too much. In some cases, the Robotics modules may expand to arbitrary equipment such as thermocyclers, tray holders, imagers, etc – that could be a future roadmap plan.
Here is some theoretical example usage below, subject to change. At this time I am deciding how much state to keep within the Perl module. By keeping state, some robot programming might be simplified (avoiding deadlock or tracking tips). In general I am aiming for a more “protocol friendly” method implementation.
To use this software with locally-connected robotics hardware:
use Robotics;
use Robotics::Tecan;
my %hardware = Robotics::query();
if ($hardware{"Tecan-Genesis"} eq "ok") {
print "Found locally-connected Tecan Genesis robotics!\n";
}
elsif ($hardware{"Tecan-Genesis"} eq "busy") {
print "Found locally-connected Tecan Genesis robotics but it is busy moving!\n";
exit -2;
}
else {
print "No robotics hardware connected\n";
exit -3;
}
my $tecan = Robotics->new("Tecan") || die;
$tecan->attach() || die; # initiate communications
$tecan->home("roma0"); # move robotics arm
$tecan->move("roma0", "platestack", "e"); # move robotics arm to vector's end
# TBD $tecan->fetch_tips($tip, $tip_rack); # move liquid handling arm to get tips
# TBD $tecan->liquid_move($aspiratevol, $dispensevol, $from, $to);
...
To use this software with remote robotics hardware over the network:
# On the local machine, run:
use Robotics;
use Robotics::Tecan;
my @connected_hardware = Robotics->query();
my $tecan = Robotics->new("Tecan") || die "no tecan found in @connected_hardware\n";
$tecan->attach() || die;
# TBD $tecan->configure("my work table configuration file") || die;
# Run the server and process commands
while (1) {
$error = $tecan->server(passwordplaintext => "0xd290"); # start the server
# Internally runs communications between client->server->robotics
if ($tecan->lastClientCommand() =~ /^shutdown/) {
last;
}
$tecan->detach(); # stop server, end robotics communciations
exit(0);
# On the remote machine (the client), run:
use Robotics;
use Robotics::Tecan;
my $server = "heavybio.dyndns.org:8080";
my $password = "0xd290";
my $tecan = Robotics->new("Tecan");
$tecan->connect($server, $mypassword) || die;
$tecan->home();
... same as first example with communication automatically routing over network ...
$tecan->detach(); # end communications
exit(0);
Some notes for those who may also want to create Perl modules for general or BioPerl use:
- Use search.cpan.org to get Module-Starter
- Run Module-Starter to create new module from module template
- Read Module::Build::Authoring
- Read Bioperl guide for authoring new modules
- Copy/write perl code into the new module
- Add POD, perl documentation
- Add unit tests into the new module
- Register for CPAN account (see CPAN wiki), register namespace
- Verify all files are in standard CPAN directory structure
- Commit & Release
Software for Biohackers
Some open source software collections of biology interest are noted here. I’ll update this list as time goes on. If you would like to have your project listed too, leave a comment with all the fields of the table and I’ll add your project. If any of these links do not work, let me know too.
| Name | Status | Field | Language | Description |
|---|---|---|---|---|
| Eclipse | Stable | Programming, editing, building, debugging | Java, C, C++, Perl, .. | Eclipse is the most widely adopted software development environment in terms of language support, corporate support, and user plugin support. It is open source. It’s the “Office” suite for programming. |
| BioPerl | Stable | Bioinformatics | Perl, C | BioPerl has many modules for genomic sequence analysis/matching, genomic searches to databases, file format conversion, etc. |
| BioPython | Stable | Bioinformatics | Python, C | BioPython has many modules for computational biology. |
| BioJava | Stable | Bioinformatics | Java | BioJava has many modules for computational biology. |
| BioLib | Stable | Bioinformatics | C, C++ | BioLib has many modules for file format conversion, integration to other Bio* language projects, genomic sequence matching, etc. |
| Bio-Linux | Stable | Operating System with Bundled Bioinformatics Applications | Many | “A dedicated bioinformatics workstation – install it or run it live” |
| DNA Linux | Stable | Operating System with Bundled Bioinformatics Applications | Many | “DNALinux is a Virtual Machine with bioinformatic software preinstalled.” |
| Several Synthetic Biology editors, simulators, or suites, listed at OpenWetWare Computational Tools, such as: Synthetic Biology Software Suite (SynBioSS), BioJADE, GenoCAD, BioStudio, BioCad,TinkerCell, Clotho |
Work In Progress | Synthetic Biology | Moslty Java, some Web based, some Microsoft .NET | Pathway modeling & simulation for synthetic biology genetic engineering, editing, parts databases, etc |
| APE (A Plasmid Editor) | Stable | Genetic engineering | Java | DNA sequence and translation editor |
“Centrifuge the column(s) at ≥10,000×g (13,000 rpm) for 1 minute, then discard the flow-through.”
A basic equation of physics, for those out there building their own centrifuges:
What are RPM, RCF, and g force and how do I convert between them?
The magnitude of the radial force generated in a centrifuge is expressed relative to the earth’s gravitational force (g force) and known as the RCF (relative centrifugal field). RCF values are denoted by a numerical number in “g” (ex. 1,000 x g). It is dependent on the speed of the rotor in revolutions per minute (RPM) and the radius of rotation. Most centrifuges are set to display RPM but have the option to change the readout to RCF.
To convert between the two by hand, use the following equation:
RCF = 11.18 (rcm) (rpm/1000)^2
Where rcm = the radius of the rotor in centimeters.
Commercial Development of Synthetic Biology Products
“BIO hosted a round-table discussion with leading-edge companies on technical and commercial advances in applications of synthetic biology. Speakers in the session represent leading firms in the field, Amyris, BioBricks Foundation, Verdezyne and Codexis.”
The Progress in Commercial Development of Synthetic Biology Applications podcast can be listened to at this link.
BIO is a biotechnology advocacy, business development and communications service organization for research and development companies in the health care, agricultural, industrial and environmental industries, including state and regional biotech associations.
Below are my notes and summary from the conference call. (Disclaimer: all quotes should be taken as terse paraphrases and see the official transcript, if any, for direct quotes.)
BIO:
“BIO sees synthetic biology as natural progression of what we’ve been doing all along [previous biology and biotech commercial research]. […] Industrial biotechnology gives us tools to selectively add genes to microbes, to allow us to engineer those microbes for the purposes of [biofuels] or production of other useful products. Synthetic biology is another tool which allows us to do this, and is an evolutionary technology, not a revolutionary technology. It grows out of what our companies have always been doing with metabolic shuffling or gene shuffling, etc. [Synthetic biology] has become so efficient that new ways of thinking about this field are necessary. We are beginning to build custom genomes from the ground up, a logical extension of the technologies [biotech companies] have developed. […] “
Industrial biotechnology’s phases:
1. Agriculture (previous phase)
2. Heathcare (previous phase)
3. and today’s phase: biofuel production, food [enrichment], environmental cleanup
Challenges in today’s world are: energy and environment (greenhouse gases, manufacturing processes, … how to also develop these in the developing world); Synthetic biology can help to address these problems.
“Every year the development times [of modifying organisms for specific tasks] are shortened [due to availability of more genomic information].”
“There is unpredictability in synthetic biology [however] this is still very manageable.”
This comment was a response to a ‘fluffy’ question about the ‘risks/dangers’ of the technology.
“[This technology is accessible because as we have heard in the news] there are now home hobbyists experimenting with this in their garage laboratories.”
Hmm; I wonder who they are talking about..
Amyris:
“We have been moving genes around for quite a while. [The difference today which yields Synthetic Biology is that] we can do things easily, rapidly and at small [measurement] scale.” Synthetic biology allows scientists to integrate all the useful [genomic, bioinformatics] data into a usable product [much more rapidly than before]. Previously it would take months to modify a microorganism, now we are down to 2-3 weeks [which is] limited only by the time required for yeast to grow [and we aren’t looking to speed that part up]; this is a rapid increase in the ability to test ideas and [measure] outputs. We view synthetic biology as very predictable [in the sense that un-intended consequences are inherently reduced]. We engineer microorganisms to grow in a [synthetic environment for fermination in a ] steel tank which reduces it’s ability to grow in a natural environment [thus] the organism loses out against environmental yeast [so modified organisms won’t cause problems in the environment since they will die]. We need more people who can understand complete pathways, complete metabolisms.”
Verdezyne:
“Synthetic Biology is a toolset to create renewable fuels and chemicals. […] The benefits of Synthetic biology are, 1. profitability, as sugar is a lower cost of carbon; 2. efficiency, from use of [standard high efficiency] fermentation processes; 3. from efficiency improvements, this improves margin, 4. decreased capital costs; 5. Use of bio-economy, using local crops [for biomass] or local photosynthetic energy to yield [chemicals for local use]. Now we can explore entire pathways in microorganisms [compared to previously when we could only look at single genes]. Traditionally, chemical engineering is the addition of chemicals to create a functionality [whereas in microbial engineering the microorganism directly creates the outputs desired]. We retooled for synthetic biology very easily [from originally building chemical engineering systems].”
Codexis:
“Biocatalysts [are] enzymes or microbes with novel properties [for commercial use]. Green alternatives to classic manufacturing routes. Biocatalysts require fewer steps and fewer harmful chemicals. Synthetic biology is one tool towards this [to] quickly create genes and pathways [using the massive amounts of genomic information now available]. [Use of] Public [genome] databases [allow us to] chop months off the [R&D] timeline. [One desire] of scientists in synthetic biology is making the microorganisms [predictable, as in in engineering] however in commercial environments we can make variants very quickly [so we can deal with variants]. There are many companies which focus on commodification of biological synthesis and we use a variety of suppliers. The analysis [the R&D] required for designing new pathways is [what is lacking in skillsets of today’s biologists].”
Drew Endy:
Patents costs are drastically more than the cost of the technology itself. The technology of the iGEM competition costs $3-4 million per year for all international teams, whereas the costs of patenting all submitted Biobricks every year would be 25k per part for 1,500 parts for a total of over $37 million dollars; thus, the patent costs are much more expensive than the technology, so this is an area which is being worked on. The next generation of biotech is hoped to “run” on an open “operating system” made from an open foundation [where new researchers can use existing genetic parts as open technology rather than having to build everything from scratch].
There was an additional analogy on the call which related synthetic biology to the emergence of vacuum tubes for electrical engineering, which ushered in incredible tools for the advancement of technology and creation of new products. I’m on the fence about these analogies, because vacuum tubes were well defined and characterized, and the shapes of their mechanical parts was well known (glass, wire, heater filaments, gas fillers, contact length, arc potentials, etc); whereas, the shapes (thus, the function) and characteristics of biological “parts” is still mostly unknown (microbiology is more than the “software strings” of nucleic acid’s A-C-G-T; it is mechanical micro-machines which interact in various ways depending on chemical context and the mechanical shapes or fittings of many of the parts are not well understood yet).
There you have it. Synthetic biology is the leaner, meaner biotech for the future.
3G Cellphone as Biotech Tool: “Cellular Phone Enabled Non-Invasive Tissue Classifier”
A recent paper in PLoS ONE describes a diagnostic system which uses a common 3G cellphone with bluetooth to assist in point-of-care measurement of tissues, from tissue samples previously taken, with remote data analysis [1]. The hope, of course, is that this could be used for detecting cancer tissue vs. non-cancer tissue. In general this technological approach is important for the following reasons: it allows data analysis across large populations with server-side storage of the data for later refinement; not all towns or cities will have expert medical staff to classify tissues at a hospital; and sending the sample to another city for classification takes time and creates measurement risk (mishandling, contamination, data entry error, biological degredation, etc). Since the tissues are measured by a digital networked device, the results can be quickly sent to a central database for further analysis, or as I hint below, for geographically mapping medical data for bioinformatics.
From my interpretation, the complete system looks like this:
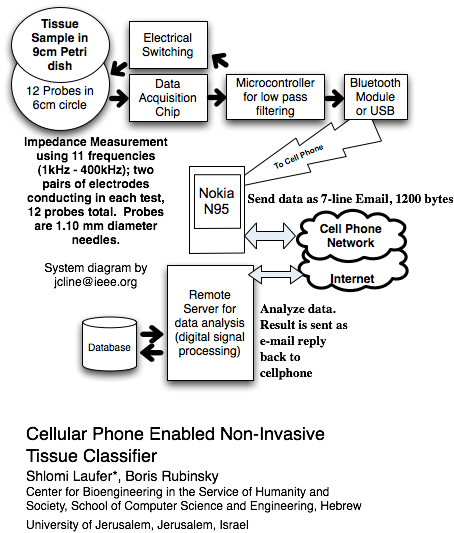
The probe electronics are described in [2]; unfortunately that article is not open access, so I can’t read it. The probes located around the sample are switched to conduct in various patterns and a learning algorithm is used to isolate the probe pair with the optimal signal. The sample is placed at the center of the petri dish and covered in saline.
Sending the raw data to a central server for analysis allows for complex pattern recognition across all samples collected; thus, the data analysis and the result can improve over time (better fitting algorithms or better weighting in the same algorithm). The impedance analysis fits according to the magnitude, phase, frequency, and the probe pair.
The article does not explain the technologies used with the cell phone for communicating between the measurement side and the cellular side (USB / Bluetooth communication link, Java, E-mail application link, etc). Though these technologies are cellphone specific, it is part of the method, and it is not described. The iPhone would be a good candidate for this project as well. A cellphone with integrated GPS would allow for location data to be sent to the server, which may be able to provide better number-crunching in the data processing algorithms, for recognition of geographic regions with high risk.
References:
[1] 2009 Cellular Phone Enabled Non-Invasive Tissue Classifier. PLoS ONE 4(4): e5178. doi:10.1371/journal.pone.0005178
[2] Ivorra A, Rubinsky B (2007) In vivo electrical impedance measurements during and after electroporation of rat liver. Bioelectrochemistry 70: 287–295.
BioMOO, the biologists’ (biohackers) virtual meeting place; in 1994
Sometime in 1994, a university obtained some funding and set up BioMOO:
BioMOO is a virtual meeting place for biologists, connected to the Globewide Network Academy. The main physical part of BioMOO is located at the BioInformatics Unit of the Weizmann Institute of Science, Israel.
BioMOO is a professional community of Biology researchers. It is a place to come meet colleagues in Biology studies and related fields and brainstorm, to hold colloquia and conferences, to explore the serious side of this new medium.
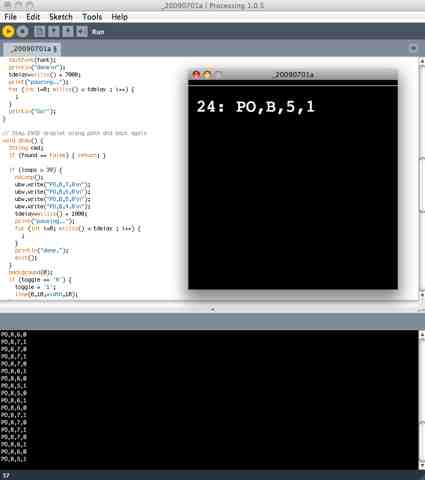
Low Cost Microcontroller-based Digital Microfluidics using “Processing”
I’ve now tested the digital microfluidics board via microcontroller. The digital microfluidics board moves a liquid droplet via Electrowetting-on-Dielectric (EWOD). The microcontroller switches the high voltage via a switching board (pictured below, using Panasonic PhotoMOS chips), which controls the +930VDC output by the HVPS (posted earlier), and runs over USB using no cost Processing.org software. This is alpha stage testing.. cleaner version to be built. The goal of course is to scale the hardware to allow automation of microbiology protocols.
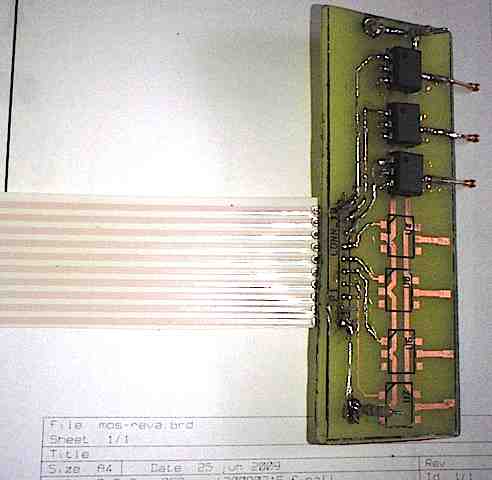

Labview is quite expensive, and industrial-grade high voltage switching boards are also quite expensive. So I built my own hardware and the Processing.org language is an easy way to test things. The Processing.org language is a free, open source graphics/media/IO layer on top of Java (as posted previously here).
What follows is the super simple test software written in Processing.org & Java.
Comments Re: Woodrow Wilson International Center’s Talk on Synthetic Biology: Feasibility of the Open Source Movement
The Woodrow Wilson International Center for Scholars hosted a recent talk on Synthetic Biology, Patents, and Open Source. This talk is now available via the web; link below. I’ve written some comments on viewing the talk, also below.
WASHINGTON – Wednesday, June 17, 2009 – Synthetic biology is developing into one of the most exciting fields in science and technology and is receiving increased attention from venture capitalists, government and university laboratories, major corporations, and startup companies. This emerging technology promises not only to enable cheap, lifesaving new drugs, but also to yield innovative biofuels that can help address the world’s energy problems.
Today, advances in synthetic biology are still largely confined to the laboratory, but it is evident from early successes that the industrial potential is high. For instance, estimates by the independent research and advisory firm Lux Research indicate that one-fifth of the chemical industry (now estimated at $1.8 trillion) could be dependent on synthetic biology by 2015.
In an attempt to enable the technology’s potential, some synthetic biologists are building their own brand of open source science. But as these researchers develop the necessary technological tools to realize synthetic biology’s promises, there is as yet no legal framework to regulate the use and ownership of the information being created.
Will this open source movement succeed? Are life sciences companies ready for open source? What level of intellectual property (IP) protection is necessary to secure industry and venture capital involvement and promote innovation? And does open source raise broader social issues? On June 17, a panel of representatives from various sectors will discuss the major challenges to future IP developments related to synthetic biology, identify key steps to addressing these challenges, and examine a number of current tensions surrounding issues of use and ownership.
________________________________
Synthetic Biology: Feasibility of the Open Source Movement
Presenters:
- Arti K. Rai, Elvin R. Latty Professor of Law, Duke Law School
- Mark Bünger, Director of Research, Lux Research
- Pat Mooney, Executive Director, ETC Group
- David Rejeski, Moderator, Director, Synthetic Biology Project
Synthetic Biology: Feasibility of the Open Source Movement
While viewing the webcast (which we are all lucky to have viewable online), I wrote some comments. Since others were interested in the comments, I’ll post ’em here.
More
Playing with the $100K Robots for Biology Automation
The Tecan Genesis Workstation 200: It’s an industrial benchtop robot for liquid handling with multiple arms for tray handling and pipetting.

The robot’s operations are complex, so an integrated development environment is used to program it (though biologists wouldn’t call it an integrated development environment; maybe they’d call it a scripting application?), with custom graphical scripting language (GUI-based) and script verification/compilation. Luckily though, the application allows third party software access and has the ability to control the robotics hardware using a minimal command set. So what to do? Hack it, of course; in this case, with Perl. This is only a headache due to Microsoft Windows incompatibilities & limitations — rarely is anything on Windows as straightforward as Unix — so as usual with Microsoft Windows software, it took about three times longer than normal to figure out Microsoft’s quirks. Give me OS/X (a real Unix) any day. Now, on to the source code!