The Woodrow Wilson International Center for Scholars hosted a recent talk on Synthetic Biology, Patents, and Open Source. This talk is now available via the web; link below. I’ve written some comments on viewing the talk, also below.
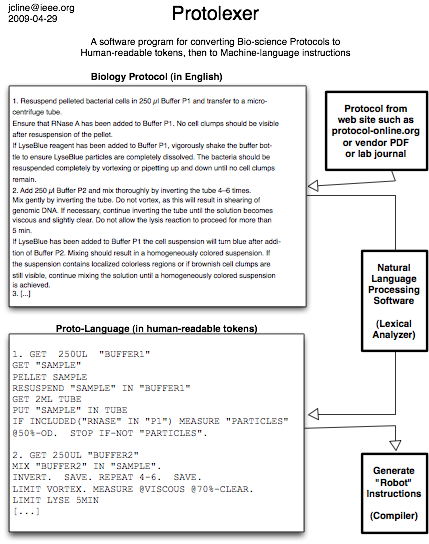
WASHINGTON – Wednesday, June 17, 2009 – Synthetic biology is developing into one of the most exciting fields in science and technology and is receiving increased attention from venture capitalists, government and university laboratories, major corporations, and startup companies. This emerging technology promises not only to enable cheap, lifesaving new drugs, but also to yield innovative biofuels that can help address the world’s energy problems.
Today, advances in synthetic biology are still largely confined to the laboratory, but it is evident from early successes that the industrial potential is high. For instance, estimates by the independent research and advisory firm Lux Research indicate that one-fifth of the chemical industry (now estimated at $1.8 trillion) could be dependent on synthetic biology by 2015.
In an attempt to enable the technology’s potential, some synthetic biologists are building their own brand of open source science. But as these researchers develop the necessary technological tools to realize synthetic biology’s promises, there is as yet no legal framework to regulate the use and ownership of the information being created.
Will this open source movement succeed? Are life sciences companies ready for open source? What level of intellectual property (IP) protection is necessary to secure industry and venture capital involvement and promote innovation? And does open source raise broader social issues? On June 17, a panel of representatives from various sectors will discuss the major challenges to future IP developments related to synthetic biology, identify key steps to addressing these challenges, and examine a number of current tensions surrounding issues of use and ownership.
________________________________
Synthetic Biology: Feasibility of the Open Source Movement
Presenters:
- Arti K. Rai, Elvin R. Latty Professor of Law, Duke Law School
- Mark Bünger, Director of Research, Lux Research
- Pat Mooney, Executive Director, ETC Group
- David Rejeski, Moderator, Director, Synthetic Biology Project
Synthetic Biology: Feasibility of the Open Source Movement
While viewing the webcast (which we are all lucky to have viewable online), I wrote some comments. Since others were interested in the comments, I’ll post ’em here.
More